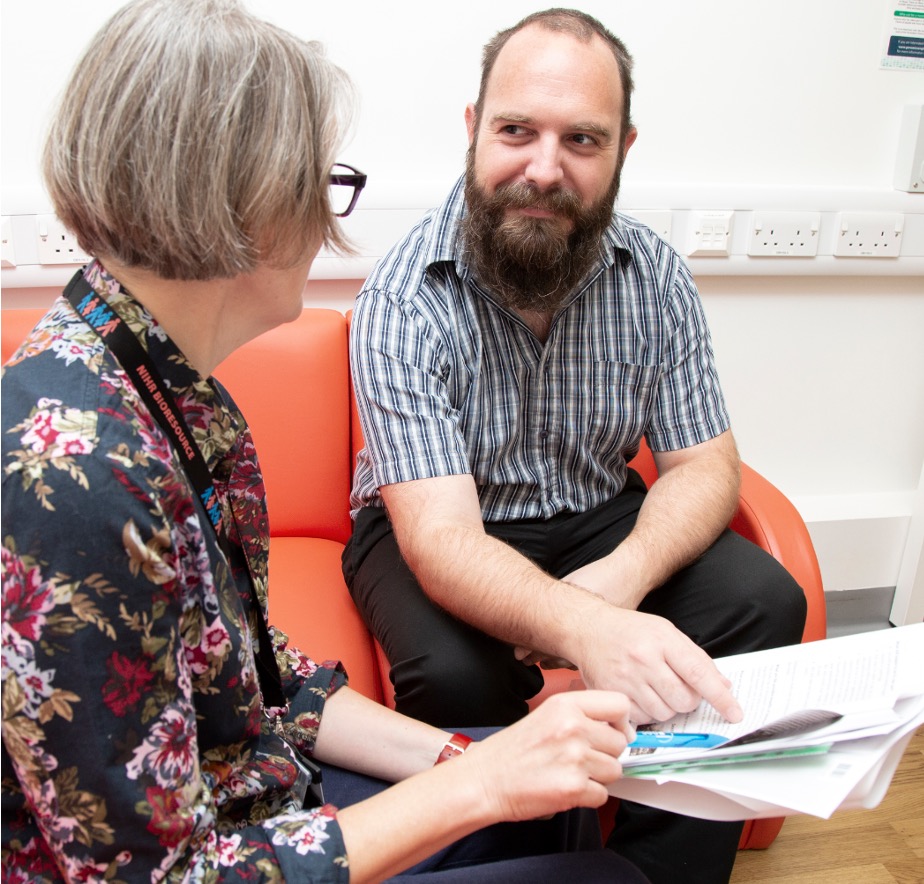
Infectious Disease Threats

Key areas of focus:
- COVID-19 vaccine responses in high-risk populations
- Detection of new viral and bacterial threats and clinical risk factors
- Characterisation of resistant bacterial and virus variants to inform treatment and vaccine design
Cambridge has a strong track record in microbiology and infectious disease research and is internationally renowned for genomic medicine. This theme aims to combine these two areas of expertise to improve detection, prevention and treatment of existing infectious diseases and help to better prepare for future pandemics.
Translating our research discoveries to the clinic or public health is at the centre of our ethos and our investigators have contributed broadly in the response to the COVID-19 pandemic. This pandemic has highlighted the need for new approaches to understand new and existing infection threats and provide a rapid pathway for new solutions.
For example, we have pioneered the use of genomics to trace outbreaks of COVID in hospitals and now are the centre of COG-UK and SARS-CoV-2 sequencing, allowing our investigators to discover how new variants emerge and their biological properties.
Our research will focus on three key areas, developing new vaccines and treatment strategies for pathogens such as SARS-CoV-2, improving the response of older people to vaccines and bolstering our response to drug-resistant bacteria.
Our approach will involve using genomic surveillance to detect and identify the viruses and other pathogens representing the biggest threats. We will be developing a data and sample repository to help us better understand the epidemiological and clinical nature of infections, and creating a resource for researchers to develop the next generation of solutions.
Here are some of the areas we are researching:
Experimental medicine for COVID-19
New vaccines and treatments for COVID-19 are emerging but a major concern is the rapid emergence of highly-resistant virus strains that could compromise global COVID-19 efforts.
It is important to understand how virus variants emerge so that we can develop strategies to mitigate against new variants and keep vulnerable immunosuppressed patients safe.
We are leading a national trial, PROTECT-V, testing the antibody sotrovimab as a prophylaxis (something which prevents the spread of disease) in more than 1,000 immune-suppressed individuals.
At the same time we are working with people living with immunosuppression across our region, to understand the challenges they and their families face regarding infectious diseases like COVID-19, and involve them in developing study protocols and patient information materials.

Evaluating and boosting COVID-19 vaccine responses in the elderly
The elderly have been largely excluded from major vaccine trials, and existing population-based studies lack age-specific detail – yet this is the group most at risk from COVID-19 and at higher risk of severe disease or death despite vaccination.
To ensure inclusion of appropriate groups, our researchers are undertaking public engagement activities in care homes and day-care centres region-wide, particularly in the more deprived areas such as Corby, where COVID-19 transmission is high, and coastal towns such as Kings Lynn which have high densities of elderly residents.
Alongside this programme, we will take the chance to explore vaccine hesitancy in the families and carers of the elderly,
and in the Traveller communities of Fenland.

Community surveillance for multi-resistant bacteria and viruses
It’s essential to track serious infections if we are to understand how they are spreading and how they might be controlled to avoid illness and death.
Through genomic surveillance (where we look at routinely collected microbiological samples, linked to hospital electronic health records) we can detect new variants and increase our understanding of how they evolve and spread.
We are working closely with hospital laboratories and NIHR BioResource to set up a system to routinely access clinical samples and organisms, on which we can perform genomic surveillance and microbiome analysis to identify key emerging bacterial and viral pathogens.
This will help us develop new therapeutic approaches for emerging pathogens including MDR (multi-drug resistant) bacteria.

