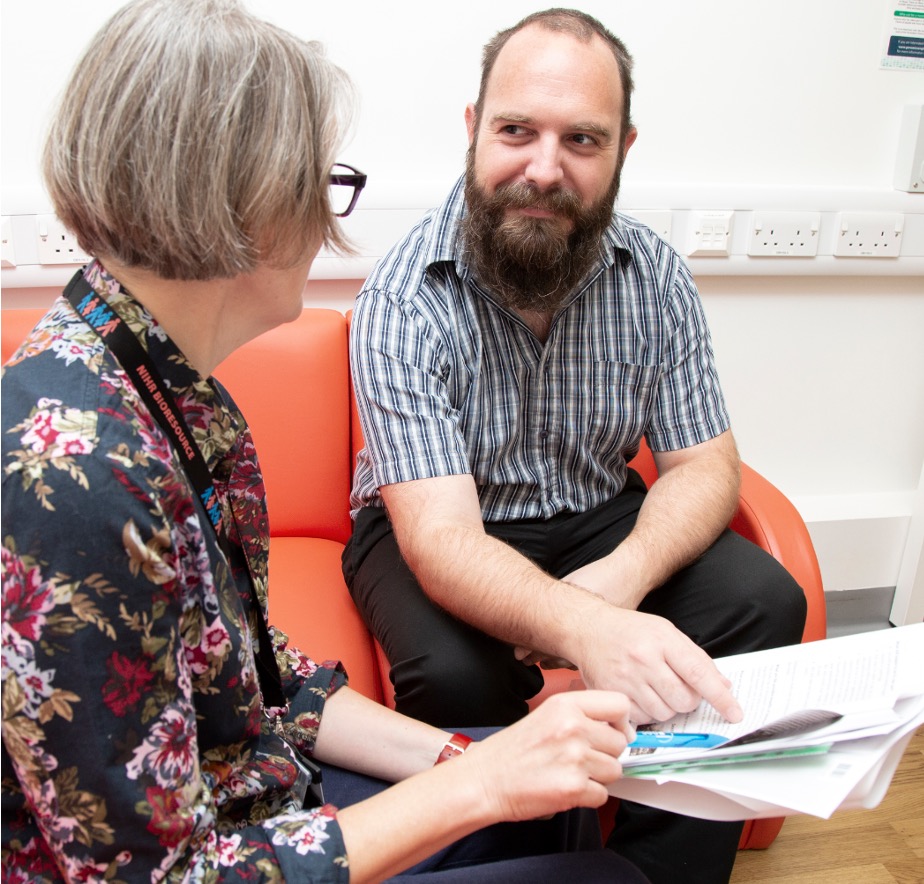
Our research: case studies
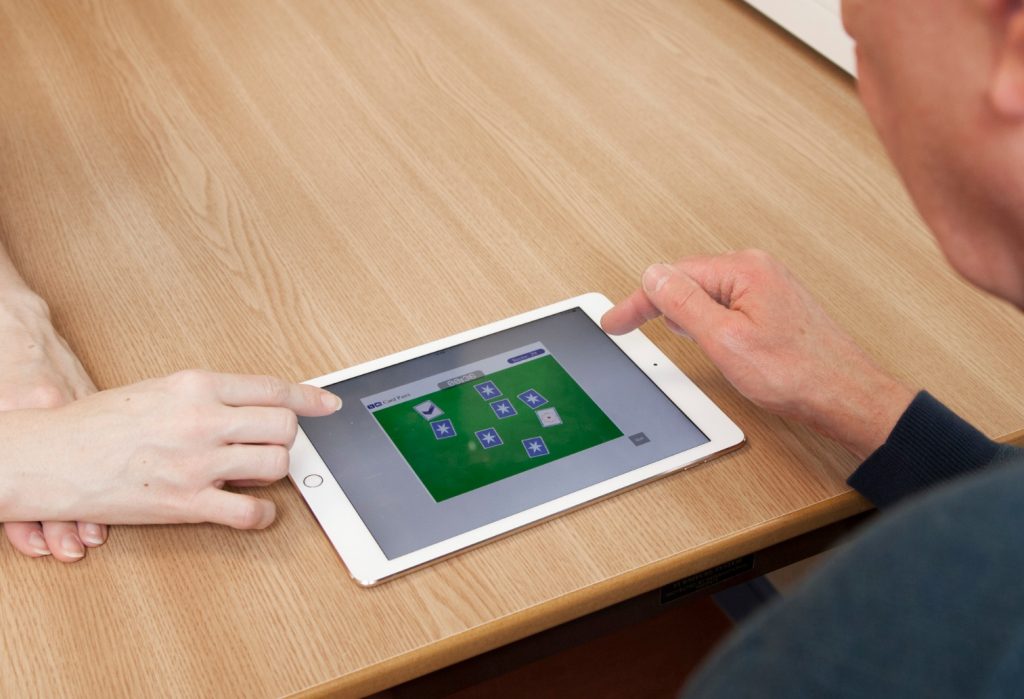

Screening for dementia in the community
Based on evidence with observational, cognitive and imaging studies, Cambridge dementia researchers have introduced community-based, computerised cognitive assessments for screening in GP surgeries.
CANTAB Mobile, which runs on an iPad, includes a learning and memory test as part of a cognitive trial. Researchers are using these tests in GP settings to detect early stages of Alzheimer’s disease. From the data collected, researchers will determine if this kind of testing in a primary care setting can detect patients earlier, before the disease progresses, allowing doctors to act earlier and put a treatment plan in place. Early therapeutic interventions are beneficial as they allow patients to stay in their homes for longer and delay institutionalised care. They also enhance quality of life for people with dementia and their carers.

Could Virtual Reality help detect the earliest stages of Alzheimer’s disease?
Researchers at the University of Cambridge, supported by the NIHR Cambridge Biomedical Research Centre (BRC), have begun a large-scale study to see if virtual reality (VR) headsets can help spot Alzheimer’s Disease years or even decades before the onset of symptoms.
The three-year project – led by Dr Dennis Chan at the University with joint funding from the Alzheimer’s Society, Merck and NIHR Cambridge BRC – will scale up earlier research in which Dr Chan’s team showed that a VR test of spatial navigation was more effective at identifying patients with mild cognitive impairment (MCI) due to Alzheimer’s disease (AD) than gold standard tests of memory and thinking currently used in clinic and research studies.
Participants (like the one photographed, above) were asked to navigate within virtual environments, displayed in state-of-the-art immersive VR headsets. This involved walking to, then remembering, the location of cones displayed within the environment. (See above image for an example: clicking on the image will open a short demonstration video.)
Now Dr Chan’s team would like to see if this test can detect AD in even earlier stages, before the onset of cognitive impairment.
In this latest study, researchers will ask 300 volunteers aged between 40 and 60, without any memory impairment, to take part in the VR study.
The volunteers have already enrolled into the PREVENT dementia programme, and have differing risks of developing AD depending on whether they have a gene that puts them at risk of AD or if they have a family history of AD.
Dr Chan’s prediction is that those at higher risk of future AD will perform less well on the VR navigation task.
Coco Newton, a PhD student on Dr Chan’s team, said: “Alzheimer’s disease is notoriously hard to detect in the early stages, when it is known as ‘preclinical’ AD. But if we can show that a VR-based navigation test is more accurate than traditional memory tests in diagnosing the preclinical disease, then that opens up lots of exciting opportunities.
“Not least it will allow people to determine whether treatments are more effective if they are started earlier and if this can delay the onset of dementia.
“By diagnosing patients much sooner we will also be able to offer them the opportunity to take part in clinical trials of future treatments.”
Testing navigation – virtually
The headset used in the study will be familiar to computer gamers.
Ms Newton explained: “You just place the headset on your face like a pair of goggles, and you are immediately placed in your ‘virtual’ environment. The headset displays images that are then used to test navigational skills as you walk around. Ultimately we want to find out if those who do worse in the tests will develop AD in later life.”
Find out more: Join Dementia Research in 3 easy steps
Can looking at the brains communication channels help detect dementia earlier?
Our brains are able to communicate with the rest of our body using connections called synapses. This is a small gap at the end of a neuron that allows a signal to pass from one neuron to the next.
These connect to neurons in the brain and to the neurons in the rest of the body. Loss of these synapses is common in early dementia.
By using a PET (positron emission tomography) scan, researchers are now able to measure the amount of synapses in the brain.
Looking at healthy volunteers who are at risk of developing dementia because of a mutation in a gene called C9orf72, they found synapse loss was already present many years before symptoms were expected, especially in a part of the brain called the thalamus.
Spotting these changes early could be vital to help those who are at high risk of dementia. Patients will be able to be monitored and begin treatment sooner.
Neuroinflammation predicts disease progression in PSP
Researchers have been able to show that Progressive Supranuclear Palsy (PSP) is associated with brain inflammation in addition to tau protein.
Progressive supranuclear palsy (PSP) is a rare neurodegenerative disease that can cause problems with balance, movement, vision, speech and swallowing. It occurs when brain cells become damaged from the build-up of protein called tau.
Patients with PSP volunteered for a trial which would involve having their brains scanned with a PET (Positron Emission Tomography) scan. The patients were then followed up for a few years.
Researchers were able to map where the inflammation occurred in the brain and highlight any significant changes. It could mean that using PET scans may be able to help clinicians see how PSP is progressing in their patients.
Read the full paper from March 21.

Studying brain activity in Progressive Supranuclear Palsy
Progressive supranuclear palsy (PSP) is a rare neurodegenerative disease that can cause problems with balance, movement, vision, speech and swallowing.
PSP is caused by the cells in the brain becoming damaged as a result of a build-up of a protein called tau.
Researchers looked at the brain activity patterns and found patients with PSP spend more time than individuals without the disease in certain brain states, meaning their brain activity was less flexible and less efficient than normal. The time spent in these brain states was more noticeable in participants who were more severely affected with PSP.
Researchers noticed that it wasn’t just certain parts of the brain that are most affected but it could affect the whole brain, even where it may appear normal on a scan or have no tau pathology.
Read the full paper published in July 21.

